|
6/13/2023 0 Comments Dna sequencing 4peaks 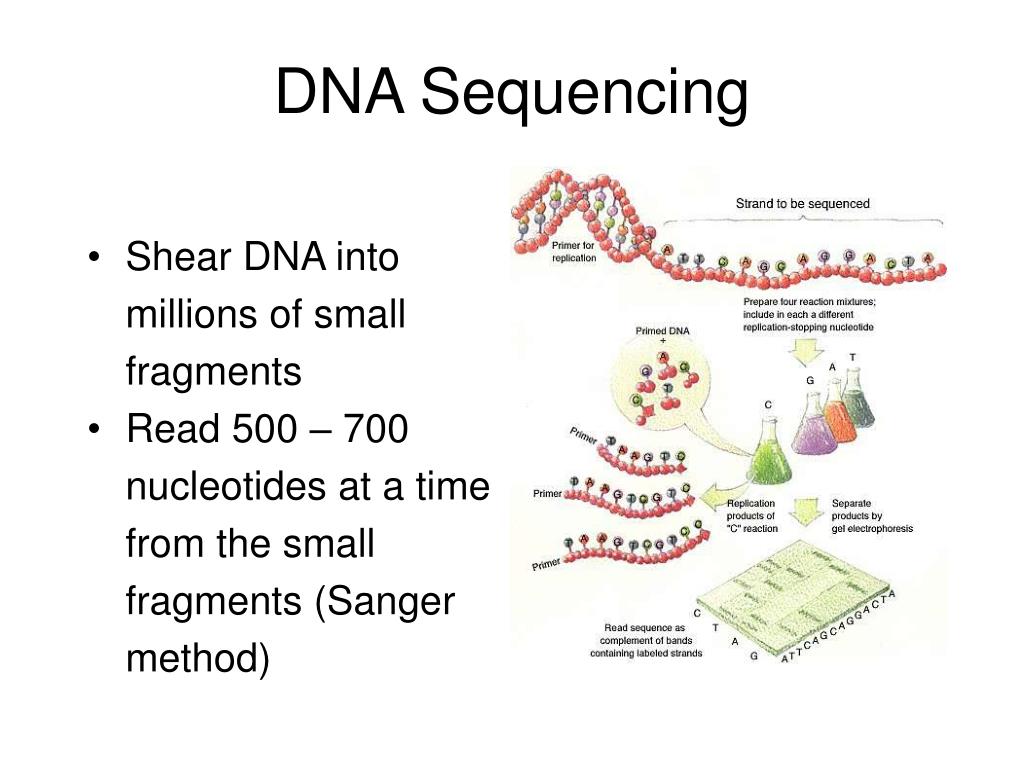
For example, mosquito host feeding patterns vary temporally, which correspond to changes in vertebrate availability and potentially the transmission of mosquito-borne viruses 1, 2. Enhanced understanding of vector-host-parasite networks may allow for integrated vector management programs focusing on highly-utilized and highly-infected host species.ĭetermining bloodmeal sources of arthropod vectors can reveal vector-host interactions critical to understanding networks of pathogen transmission and guiding vector control and disease prevention campaigns. Up to four bloodmeal sources were detected in a single triatomine ( I. The amplicon deep sequencing results (average of 302,080 usable reads per sample) replicated the host community revealed using Sanger sequencing, and detected additional hosts in five triatomines (13.9%), including two additional blood sources ( Procyon lotor and Bassariscus astutus). Samples sequenced by the Sanger method were also subjected to Illumina MiSeq amplicon deep sequencing. Sanger sequencing revealed 15 distinct host species, which included humans, domestic animals ( Canis lupus familiaris, Ovis aries, Gallus gallus, Bos taurus, Felis catus, and Capra hircus), wildlife ( Rattus rattus, Incilius nebulifer, Sciurus carolinensis, Sciurus niger, and Odocoileus virginianus), and captive animals ( Panthera tigris, Colobus spp., and Chelonoidis carbonaria). Direct Sanger sequencing of PCR amplicons was successful for 36 samples (31%). We used two methods-Sanger sequencing and amplicon deep sequencing-to target a 228 bp region of the vertebrate Cytochrome b gene and determine hosts fed upon by triatomines (n = 115) collected primarily in Texas, USA. 
However, the ability of methods to distinguish bloodmeal sources varies widely. Knowledge of host associations of blood-feeding vectors may afford insights into managing disease systems and protecting public health. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
